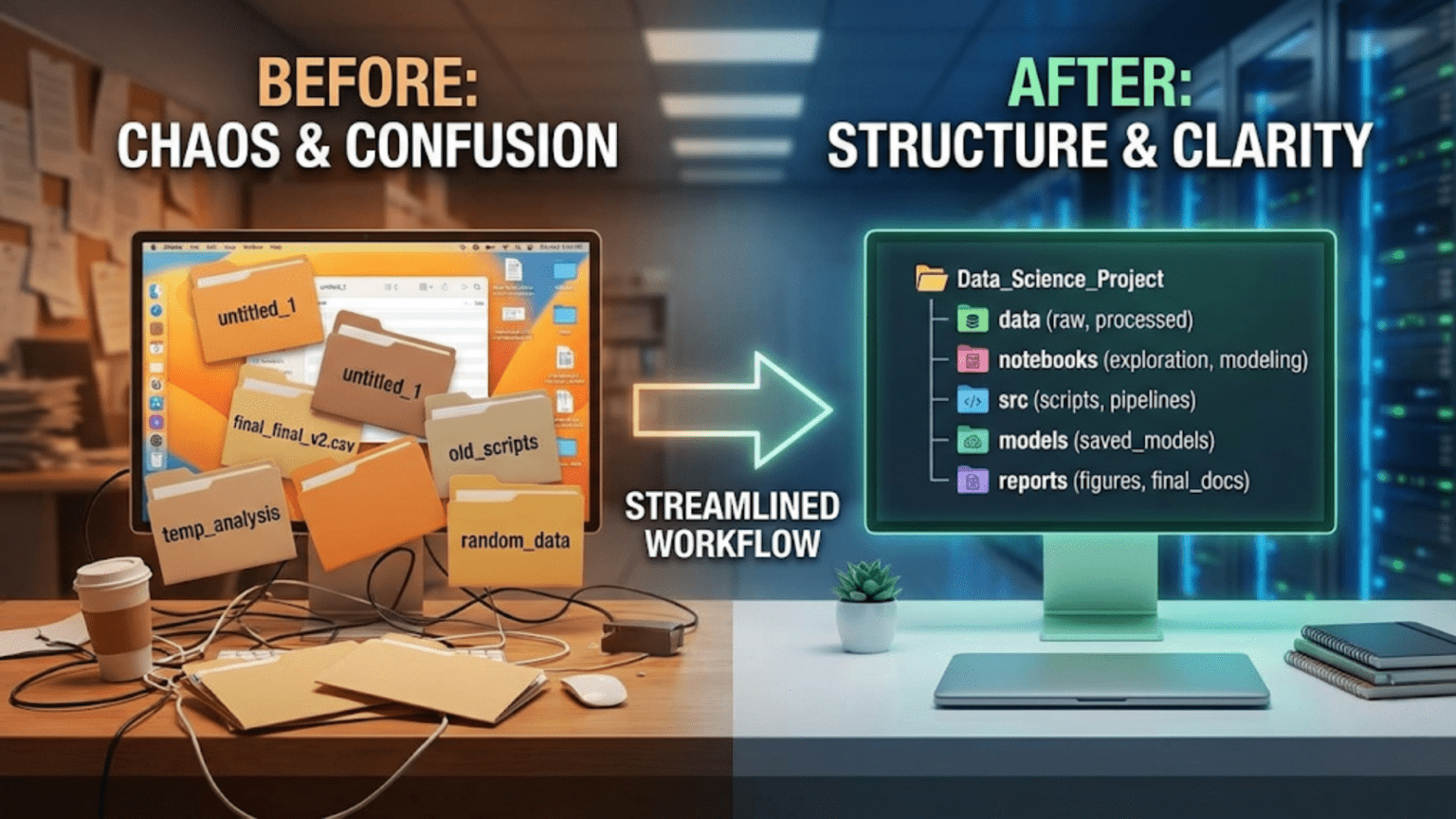
A well-organized data science project uses a consistent, logical folder structure that separates raw data from processed data, exploratory notebooks from production code, and configuration from source logic — making the project easy to navigate, reproduce, and hand off to teammates. The most widely adopted standard is the Cookiecutter Data Science template, which provides a battle-tested directory structure used by data science teams at organizations worldwide.
Introduction
Imagine opening a data science project you worked on eight months ago. You need to reproduce a model result for a stakeholder presentation. Where is the data? Which notebook has the final version of the feature engineering? Is model_v3_final_USE_THIS.pkl actually the best model, or was there a model_v4 somewhere? Why are there three files called preprocessing.py in different directories?
This scenario, embarrassingly common in data science, is entirely preventable. The difference between a project you can confidently revisit after months away and a project that’s effectively lost to time is almost entirely a matter of organization.
Unlike software engineering, where organizational patterns like MVC (Model-View-Controller) have been standardized for decades, data science is a younger discipline that evolved partly from ad-hoc research workflows. Many data scientists learned their craft through Kaggle notebooks, academic papers, and self-directed tutorials — all of which prioritize getting answers fast over building maintainable project structures.
The good news is that the data science community has developed strong organizational standards over the past decade. These standards, refined through hard experience at companies ranging from small startups to major tech firms, strike a careful balance: structured enough to be navigable and reproducible, flexible enough to accommodate the messiness of real data science work.
This guide covers everything you need to organize your data science projects like a professional: from the fundamental principles behind good organization, through specific directory structures and naming conventions, to templates and tools that make the right structure the default rather than an extra effort.
Why Project Organization Matters
Before diving into how to organize, it’s worth establishing why organization matters so deeply in data science specifically — because the stakes are higher here than in many other domains.
Reproducibility
Data science is fundamentally about producing results that can be trusted and verified. A model’s performance claim means nothing if no one can reproduce it. Good project organization means that every result is traceable: which data was used, how it was preprocessed, which code version trained the model, and what parameters were set. Without this traceability, you’re producing art, not science.
The Handoff Problem
Projects change hands constantly in professional settings. You might leave a team, bring in a new data scientist to scale a project, or need a colleague to cover for you during an absence. An organized project can be understood by someone new in hours; a disorganized one requires weeks of archaeology and still leaves uncertainty.
The Return Problem
You will return to your own projects after months away. Memory is unreliable. The mental model you had while building the project will have faded. Organized projects with clear structure, naming conventions, and documentation are self-explanatory — they don’t depend on you remembering things you’ve forgotten.
Collaboration at Scale
As data science teams grow, uncoordinated individual styles create friction. When five team members have five different folder structures, every new project requires everyone to relearn navigation. A shared standard means anyone can immediately find what they need in any team member’s project.
Debugging and Iteration
Data science involves a lot of iteration — trying different models, different features, different preprocessing approaches. Without structure, iteration creates an accumulating pile of half-finished experiments with unclear relationships. With structure, each experiment is clearly contextualized: what was tried, when, with what result.
Core Principles of Good Data Science Project Organization
Before specific structures and templates, some universal principles that should guide all organizational decisions.
Principle 1: Separate Data from Code
Data and code have fundamentally different natures and should never be mixed:
- Data is often large, binary, and should not be version-controlled in Git
- Data changes on its own (upstream sources update, new data arrives)
- Data has different access controls than code (often more sensitive)
- Code generates processed data from raw data — they have a directional relationship
Keep all data in a dedicated data/ directory, with clear subdirectories for raw, processed, and intermediate states. Keep code completely separate.
Principle 2: Raw Data Is Sacred
Raw data — the original, unmodified data you received from a source — must be treated as read-only and never modified. All transformations produce new files in separate directories. This ensures you can always go back to the original source and reproduce your work from scratch.
Think of raw data like a master tape recording: you make copies to work with, but you never edit the original.
Principle 3: Separate Exploration from Production
Exploratory notebooks are messy by nature — they contain dead ends, experimental code, half-written analyses, and commented-out approaches you might return to. Production code must be clean, tested, and reliable. These two categories need separate homes and different standards of care.
Principle 4: Make the Project Self-Documenting
The directory names, file names, and project structure itself should communicate what the project contains and how it flows. Someone who has never seen the project should be able to understand its general shape by browsing the file tree for two minutes.
Principle 5: Automate the Boring Parts
Configuration, paths, environment setup, and pipeline execution should all be codified so they require no manual intervention or memorization. A Makefile, a configuration file, and a well-written README eliminate the hidden tribal knowledge that makes projects fragile.
The Standard Data Science Project Structure
This is the directory structure used — with variations — by professional data science teams worldwide. It’s informed by the Cookiecutter Data Science template (drivendata.github.io/cookiecutter-data-science), which is itself the distillation of industry best practices.
project-name/
│
├── data/
│ ├── raw/ ← Original, immutable source data
│ ├── external/ ← Data from external sources (third-party)
│ ├── interim/ ← Partially processed, intermediate data
│ └── processed/ ← Final, clean data ready for modeling
│
├── notebooks/
│ ├── exploratory/ ← Messy EDA, experiments, dead ends — anything goes
│ │ ├── 01_eda_initial.ipynb
│ │ ├── 02_feature_ideas.ipynb
│ │ └── 03_model_experiments.ipynb
│ └── reports/ ← Clean, polished notebooks for sharing results
│ └── 01_final_analysis.ipynb
│
├── src/ ← Source code (Python modules)
│ ├── __init__.py
│ ├── data/
│ │ ├── __init__.py
│ │ ├── make_dataset.py ← Scripts to download or generate data
│ │ └── preprocess.py ← Data cleaning and transformation
│ ├── features/
│ │ ├── __init__.py
│ │ └── build_features.py ← Feature engineering
│ ├── models/
│ │ ├── __init__.py
│ │ ├── train_model.py ← Model training
│ │ ├── predict_model.py ← Model inference
│ │ └── evaluate_model.py ← Model evaluation
│ └── visualization/
│ ├── __init__.py
│ └── visualize.py ← Reusable visualization functions
│
├── models/ ← Trained model files (often gitignored)
│ ├── baseline_lr.pkl
│ └── best_model_xgb.pkl
│
├── reports/ ← Generated analysis and output documents
│ ├── figures/ ← Saved plots and visualizations
│ │ ├── feature_importance.png
│ │ └── confusion_matrix.png
│ └── final_report.pdf
│
├── tests/ ← Unit and integration tests
│ ├── __init__.py
│ ├── test_preprocess.py
│ ├── test_features.py
│ └── test_models.py
│
├── configs/ ← Configuration files
│ ├── config.yaml ← Main project configuration
│ └── model_config.yaml ← Model hyperparameters
│
├── .github/
│ └── workflows/
│ └── tests.yml ← CI/CD pipeline
│
├── .gitignore ← Files excluded from version control
├── .env.example ← Template for environment variables (no secrets)
├── Makefile ← Automation commands
├── README.md ← Project documentation
├── requirements.txt ← Production dependencies
├── requirements-dev.txt ← Development + testing dependencies
└── pyproject.toml ← Tool configuration (black, pytest, etc.)Let’s go through each directory in detail.
Directory Deep Dive
The data/ Directory
This is one of the most important directories to get right.
data/
├── raw/ ← Never modified after receipt. Gitignored.
├── external/ ← Third-party data: Census data, market data, etc.
├── interim/ ← Output of partial processing (e.g., after deduplication)
└── processed/ ← Final, model-ready data. Gitignored.Key rules:
data/raw/is read-only. No script ever writes to it. It represents the ground truth of what you received.- Every subdirectory is typically gitignored (data files are too large and too sensitive for Git). Use cloud storage (S3, GCS, Azure Blob) or DVC for data version control.
- Add a
data/raw/README.mddescribing where the data came from, when it was collected, what format it’s in, and any known issues. This readme is committed to Git even though the data files aren’t.
# data/raw/README.md
## Customer Transaction Data
**Source**: Internal CRM database export
**Date collected**: 2024-09-15
**Format**: CSV
**File**: transactions_2024_q3.csv (412 MB, gitignored)
**Description**: All customer transactions for Q3 2024.
**Schema**: customer_id (str), transaction_date (datetime), amount (float),
product_id (str), channel (str)
**Known issues**: ~2% of records have null values in 'channel' column.
**Access**: Available in S3 bucket s3://company-data/raw/transactions/This README costs 10 minutes to write and saves hours of confusion later.
The notebooks/ Directory
notebooks/
├── exploratory/ ← Personal lab notebooks, messy is fine
└── reports/ ← Polished notebooks for sharingNotebook naming convention: Number notebooks with a prefix to enforce chronological ordering and include a brief description:
01_initial_data_exploration.ipynb
02_missing_value_analysis.ipynb
03_feature_correlation_study.ipynb
04_baseline_model_experiments.ipynb
05_xgboost_hyperparameter_tuning.ipynbThis naming gives you a natural narrative: someone new to the project can read notebooks in order and follow the analytical journey. The number prefix also keeps them sorted correctly in file explorers.
The split between exploratory/ and reports/ is important. Exploratory notebooks are your personal lab: messy, iterative, containing dead ends, debug output, and commented-out experiments. They don’t need to run cleanly from top to bottom — they’re your thinking made visible.
Report notebooks are cleaned-up presentations of findings. They should run cleanly from Kernel → Restart and Run All, contain Markdown explanations of each section, and be suitable for sharing with stakeholders or committing as project documentation.
A practical rule: exploratory notebooks are for you; report notebooks are for everyone else.
The src/ Directory
This is where your reusable, production-quality Python code lives. As you work in notebooks and discover patterns and functions worth keeping, you extract them here.
src/
├── __init__.py
├── data/
│ ├── make_dataset.py ← Download, generate, or receive raw data
│ └── preprocess.py ← Clean and transform raw data
├── features/
│ └── build_features.py ← Create model features from processed data
├── models/
│ ├── train_model.py ← Training logic
│ ├── predict_model.py ← Inference logic
│ └── evaluate_model.py ← Metrics and evaluation
└── visualization/
└── visualize.py ← Reusable plot functionsThe __init__.py files make each directory a Python package, enabling clean imports:
# In a notebook or script:
from src.data.preprocess import clean_transactions
from src.features.build_features import add_customer_lifetime_value
from src.models.train_model import train_xgboostThis import pattern means your notebooks stay clean: they orchestrate functions rather than containing all the logic inline.
A well-developed src/ directory is the hallmark of a mature data science project. It represents the crystallization of exploratory work into a stable, tested codebase.
The configs/ Directory
Hardcoding values like file paths, model hyperparameters, and column names throughout your code is a maintenance nightmare. Config files centralize all these values:
# configs/config.yaml
paths:
raw_data: "data/raw/transactions_2024_q3.csv"
processed_data: "data/processed/transactions_clean.csv"
model_output: "models/best_model.pkl"
data:
target_column: "churned"
id_column: "customer_id"
date_column: "transaction_date"
categorical_columns:
- "channel"
- "product_category"
- "region"
training:
test_size: 0.2
random_state: 42
cv_folds: 5
logging:
level: "INFO"
file: "logs/training.log"# configs/model_config.yaml
xgboost:
n_estimators: 500
learning_rate: 0.05
max_depth: 6
min_child_weight: 3
subsample: 0.8
colsample_bytree: 0.8
gamma: 0.1
reg_alpha: 0.1
reg_lambda: 1.0
random_state: 42
random_forest:
n_estimators: 300
max_depth: 10
min_samples_split: 5
min_samples_leaf: 2
random_state: 42Load configs in your Python code with PyYAML:
import yaml
from pathlib import Path
def load_config(config_path: str = "configs/config.yaml") -> dict:
with open(config_path, 'r') as f:
return yaml.safe_load(f)
config = load_config()
raw_data_path = config['paths']['raw_data']
target_col = config['data']['target_column']Now changing a file path or hyperparameter requires editing one YAML file, not hunting through Python code across multiple modules.
The tests/ Directory
Tests are frequently skipped in data science projects. This is a mistake that compounds over time — untested code is fragile code, and fragile code becomes impossible to refactor.
tests/
├── __init__.py
├── test_preprocess.py
├── test_features.py
└── test_models.pyTests don’t need to be exhaustive to be valuable. Even a handful of tests for your most critical functions provides enormous value:
# tests/test_preprocess.py
import pandas as pd
import pytest
from src.data.preprocess import clean_transactions, handle_missing_values
class TestCleanTransactions:
def test_removes_duplicate_rows(self):
df = pd.DataFrame({
'customer_id': ['A', 'A', 'B'],
'amount': [100.0, 100.0, 200.0]
})
result = clean_transactions(df)
assert len(result) == 2
def test_preserves_all_columns(self):
df = pd.DataFrame({
'customer_id': ['A', 'B'],
'amount': [100.0, 200.0],
'channel': ['web', 'mobile']
})
result = clean_transactions(df)
assert set(result.columns) == set(df.columns)
def test_handles_empty_dataframe(self):
df = pd.DataFrame()
result = clean_transactions(df)
assert len(result) == 0
class TestHandleMissingValues:
def test_fills_numeric_with_median(self):
df = pd.DataFrame({'amount': [100.0, None, 300.0]})
result = handle_missing_values(df)
assert result['amount'].isnull().sum() == 0
assert result['amount'].iloc[1] == 200.0 # Median of 100, 300Run with: pytest tests/ -v
The Makefile
A Makefile codifies common project commands into named, executable targets — eliminating “how do I run this again?” questions:
# Makefile
.PHONY: env data features train evaluate test lint clean
# Set up the development environment
env:
python -m venv venv
source venv/bin/activate && pip install -r requirements-dev.txt
@echo "Environment ready. Run 'source venv/bin/activate' to activate."
# Download and prepare raw data
data:
python src/data/make_dataset.py --config configs/config.yaml
# Build features from processed data
features:
python src/features/build_features.py --config configs/config.yaml
# Train the model
train:
python src/models/train_model.py --config configs/config.yaml
# Evaluate the model
evaluate:
python src/models/evaluate_model.py --config configs/config.yaml
# Run the full pipeline
pipeline: data features train evaluate
@echo "Full pipeline complete."
# Run tests
test:
pytest tests/ -v --tb=short
# Run linting and formatting checks
lint:
black --check src/ tests/
flake8 src/ tests/
# Format code
format:
black src/ tests/
isort src/ tests/
# Clean generated files
clean:
find . -type f -name "*.pyc" -delete
find . -type d -name "__pycache__" -exec rm -rf {} +
find . -name ".ipynb_checkpoints" -exec rm -rf {} +With this Makefile, a new team member can run make env to set up their environment, make pipeline to run the entire data pipeline, and make test to run tests — without reading any documentation about which scripts to run in which order.
File Naming Conventions
Consistent naming conventions make file navigation fast and intuitive. Here are the standards used by professional data science teams.
General Rules
- Use lowercase with underscores for all filenames:
feature_engineering.pynotFeatureEngineering.pyorfeature-engineering.py - Be descriptive but concise:
train_xgboost_classifier.pynottrain.pyorthis_is_the_training_script_for_the_xgboost_model_v2.py - Avoid version numbers in filenames: use Git for versioning, not
model_v3_final_FINAL.pkl - Use ISO date format when dates are needed:
data_20240915.csvnotdata_Sept_15.csv
Notebook Naming
# Pattern: NN_brief_description.ipynb
# NN = two-digit number for ordering
01_data_exploration.ipynb
02_missing_value_analysis.ipynb
03_feature_engineering_experiments.ipynb
04_baseline_model_comparison.ipynb
05_xgboost_optimization.ipynb
06_final_model_evaluation.ipynbScript Naming
Use the verb-noun pattern for scripts that perform actions:
# Good — verb describes what the script does
make_dataset.py
build_features.py
train_model.py
evaluate_model.py
generate_report.py
clean_transactions.py
# Avoid — too generic
main.py (unless it truly is a main entry point)
utils.py (what utilities? be specific)
helpers.py (helpers for what?)
script.pyData File Naming
# Pattern: description_YYYYMMDD.extension or description_version.extension
transactions_raw_20240915.csv ← Raw data with acquisition date
customers_processed_v2.parquet ← Processed data with version
features_train_set.parquet ← Role-specific (train/test)
features_test_set.parquetModel File Naming
# Pattern: algorithm_experiment_description.extension
baseline_logistic_regression.pkl
xgboost_v1_default_params.pkl
xgboost_v2_tuned_lr0.05_depth6.pkl
best_model_xgboost.pkl ← Clearly marked best performerAdapting the Structure for Different Project Types
The standard structure works well for most projects, but different problem types call for slight adaptations.
Small Exploratory Projects
For quick, one-person analyses that won’t be deployed or shared widely, a lighter structure is appropriate:
quick-analysis/
├── data/
│ └── raw/
├── notebooks/
│ └── 01_analysis.ipynb
├── .gitignore
├── requirements.txt
└── README.mdDon’t over-engineer short-lived work. The full structure shines for projects that will grow, be shared, or need to be maintained.
Machine Learning Pipeline Projects
For projects that deploy models into production:
ml-pipeline/
├── data/ ← (standard)
├── src/
│ ├── data/ ← (standard)
│ ├── features/ ← (standard)
│ ├── models/ ← (standard)
│ └── api/ ← NEW: model serving code
│ ├── app.py ← Flask/FastAPI application
│ ├── schemas.py ← Request/response schemas
│ └── predict.py ← Prediction endpoint logic
├── notebooks/ ← (standard)
├── tests/ ← (standard)
├── configs/ ← (standard)
├── docker/ ← NEW: containerization
│ ├── Dockerfile
│ └── docker-compose.yml
├── deployment/ ← NEW: infrastructure as code
│ ├── kubernetes/
│ └── terraform/
└── monitoring/ ← NEW: model monitoring
└── drift_detection.pyNLP Projects
Natural language processing projects add specialized data directories:
nlp-project/
├── data/
│ ├── raw/ ← Original documents, corpora
│ ├── interim/
│ │ └── tokenized/ ← Tokenized text
│ └── processed/
│ └── embeddings/ ← Vector representations
├── src/
│ ├── data/
│ │ ├── scraper.py ← Data collection scripts
│ │ └── tokenizer.py ← Text tokenization
│ ├── features/
│ │ └── text_features.py ← TF-IDF, embeddings, etc.
│ └── models/
│ ├── fine_tune.py ← Model fine-tuning
│ └── evaluate.py
├── models/
│ └── fine_tuned/ ← Fine-tuned model checkpoints
└── notebooks/Computer Vision Projects
cv-project/
├── data/
│ ├── raw/
│ │ ├── images/ ← Original images
│ │ └── annotations/ ← Labels, masks, bounding boxes
│ └── processed/
│ └── augmented/ ← Augmented training images
├── src/
│ ├── data/
│ │ ├── dataset.py ← PyTorch/TF Dataset class
│ │ └── augmentation.py ← Data augmentation pipeline
│ └── models/
│ ├── architecture.py ← Neural network architecture
│ └── train.py
├── models/
│ └── checkpoints/ ← Training checkpoints
└── notebooks/The Cookiecutter Data Science Template
Rather than setting up this structure from scratch each time, use the Cookiecutter Data Science template to generate it automatically.
Using Cookiecutter
# Install cookiecutter
pip install cookiecutter
# Generate a new project from the template
cookiecutter https://github.com/drivendata/cookiecutter-data-scienceCookiecutter asks a series of questions and generates the complete directory structure:
project_name [project_name]: customer-churn-model
repo_name [customer-churn-model]: customer-churn-model
author_name [Your name]: Jane Smith
description [A short description of the project.]: Predict customer churn using transaction history
open_source_license [MIT]: MIT
python_interpreter [python3]: python3The generated structure is ready to use immediately, with placeholder README files in each directory explaining its purpose.
Customizing the Template
If your team has specific needs, you can fork the Cookiecutter Data Science template and add your organization’s standard files: company-specific .gitignore additions, your team’s Makefile targets, standard CI/CD configurations, and custom documentation templates. Every new project then starts from your organization’s opinionated standard.
Managing the Data Directory with DVC
Since data files are typically gitignored, you need a separate system for data version control. DVC (Data Version Control) extends Git-like semantics to data and model files.
DVC Basics
# Install DVC
pip install dvc
# Initialize DVC in your Git repository
dvc init
# Add a data file to DVC tracking
dvc add data/raw/transactions_2024_q3.csv
# Creates: data/raw/transactions_2024_q3.csv.dvc (small metadata file)
# The .dvc file IS committed to Git
# The actual CSV is NOT committed (DVC manages it separately)
# Configure remote storage (e.g., S3)
dvc remote add -d myremote s3://my-bucket/dvc-store
# Push data to remote storage
dvc push
# Teammates pull data
dvc pullWith DVC, your data/raw/transactions_2024_q3.csv.dvc file is committed to Git and records the exact version of the data (via a hash). Teammates run dvc pull and get the exact same data file. The data lives in S3 (or GCS, Azure, or even a shared network drive), not in Git.
This gives you data version control that parallels code version control — you can dvc checkout to get the data that corresponds to any Git commit, making complete experiment reproducibility possible.
Documentation Within the Project
Good project organization isn’t just about directory structure — it’s about the documentation embedded throughout the project.
The README.md
The project-level README should cover:
# Customer Churn Prediction
Predict which customers are likely to churn in the next 30 days
using transaction history and behavioral features.
## Project Organization
├── data/
│ ├── raw/ ← Original transaction data (DVC-managed)
│ └── processed/ ← Cleaned, feature-engineered data
├── notebooks/ ← Exploratory and report notebooks
├── src/ ← Python source modules
├── tests/ ← Unit tests
├── configs/ ← Configuration files
└── models/ ← Trained model artifacts
## Setup
Prerequisites: Python 3.11, DVC
git clone git@github.com:team/customer-churn.git
cd customer-churn
make env
source venv/bin/activate
dvc pull # Download data from remote storage
## Running the Pipeline
make pipeline # Run complete data → features → train → evaluate pipeline
## Results
| Model | AUC-ROC | F1 Score | Precision | Recall |
|---------------|---------|----------|-----------|--------|
| Baseline (LR) | 0.823 | 0.764 | 0.771 | 0.757 |
| Random Forest | 0.891 | 0.819 | 0.834 | 0.804 |
| **XGBoost** | **0.914** | **0.847** | **0.856** | **0.839** |
Best model: XGBoost (see `notebooks/reports/06_final_model_evaluation.ipynb`)README Files in Subdirectories
Each major subdirectory should have its own README explaining its contents:
# data/raw/README.md
Source: CRM database export, Q3 2024.
Files: transactions_2024_q3.csv (DVC-tracked). See .dvc file.
Schema: customer_id, transaction_date, amount, product_id, channel.
Access: Request access via s3://company-data/raw/ (ask DevOps team).# models/README.md
Models are saved as pickle files with scikit-learn/XGBoost.
Naming: algorithm_experiment_description.pkl
Best production model: best_model_xgboost.pkl (AUC-ROC: 0.914)
Training details: notebooks/reports/06_final_model_evaluation.ipynbThese in-directory READMEs are small time investments that make the project self-navigating.
Common Anti-Patterns to Avoid
Recognizing bad patterns is as important as knowing good ones.
Anti-Pattern 1: Everything in One Directory
project/ ← All files dumped here
├── analysis.ipynb
├── analysis_v2.ipynb
├── analysis_final.ipynb
├── data.csv
├── clean_data.csv
├── model.pkl
├── model_final.pkl
├── preprocessing.py
├── train.py
└── utils.pyNo separation of concerns, no navigability, no reproducibility. A project archaeologist’s nightmare.
Anti-Pattern 2: Version Numbers in Filenames
model_v1.pkl
model_v2.pkl
model_v2_improved.pkl
model_v3_FINAL.pkl
model_v3_FINAL_fixed.pkl
model_v3_FINAL_fixed_USE_THIS.pklThis is what Git is for. Use one best_model.pkl and let commit history tell the story of how you got there.
Anti-Pattern 3: Hardcoded Paths
# Fragile — breaks on anyone else's machine
df = pd.read_csv("/Users/jane/Desktop/project/data/my_data.csv")
# Better — relative path using config
df = pd.read_csv(config['paths']['raw_data'])
# Also better — using pathlib
from pathlib import Path
DATA_DIR = Path(__file__).parent.parent / "data"
df = pd.read_csv(DATA_DIR / "raw" / "transactions.csv")Hardcoded absolute paths guarantee that your code runs only on your machine.
Anti-Pattern 4: Committing Data to Git
Large data files in Git repositories slow down every clone, push, and pull for everyone on the team and fill up the repository. Use DVC, Git LFS, or simply gitignore data directories and document access via README.
Anti-Pattern 5: Notebooks as Production Code
Notebooks are excellent for exploration and communication, but they shouldn’t be the operational code that runs your pipeline. Complex, multi-notebook pipelines that must be run in a specific order, manually, are brittle and non-automatable. Extract the logic into src/ modules and orchestrate them with a Makefile or pipeline tool.
Evolving Your Structure Over Time
Projects grow. A structure appropriate for day one may feel constraining on day ninety. The key is to evolve deliberately rather than letting entropy accumulate.
Signs your structure needs refactoring:
- You’re spending more than a minute finding files you know exist
- New team members ask where to put things more than once per day
- Your notebooks directory has more than 20 files with no subdirectory organization
- The same logic appears in three different places with slight variations
- Nobody knows which model file is the current production model
When to refactor:
- At natural project milestones (completing EDA, completing initial model, preparing for deployment)
- When onboarding new team members (their confusion is useful signal)
- Before sharing the project publicly or presenting it to stakeholders
Refactoring directory structure is like refactoring code — it should be done regularly in small increments, not deferred until the project is completely unmaintainable.
Summary
Organizing data science project files is not bureaucratic overhead — it is the infrastructure of reproducible, collaborative, maintainable work. The investment in a clean structure pays dividends every time you return to a project, onboard a colleague, reproduce a result, or hand off your work to a team.
The standard structure — with clearly separated data/, notebooks/, src/, tests/, configs/, and models/ directories — reflects hard-won experience about how data science projects actually work: data flows from raw to processed, exploration yields code that gets extracted into modules, modules get tested, and configuration drives behavior without hardcoding.
The practical tools that support this structure — Cookiecutter for generating it, DVC for data versioning, Makefiles for automation, config YAML files for parameterization — transform good intentions into consistent execution. Use them on every project, from the very first commit, and the organizational habits they instill will make you a significantly more effective data scientist.
Key Takeaways
- Good project organization enables reproducibility, collaboration, and the ability to return to projects months later without confusion — it is not optional overhead but a fundamental professional practice
- Raw data is sacred and read-only; all processing produces new files in separate directories, ensuring you can always reproduce work from the original source
- The standard structure separates data from code, exploration from production, and configuration from logic — each separation solves a distinct category of organizational problem
- Notebook naming with numeric prefixes (
01_eda.ipynb,02_features.ipynb) creates a navigable narrative of the analytical process - Config YAML files eliminate hardcoded values and make experiments parameterizable — one file to change file paths and hyperparameters across the entire project
- A
Makefilecodifies common commands so anyone can runmake trainormake testwithout reading documentation about which scripts to run in which order - DVC (Data Version Control) extends Git semantics to large data and model files, making complete experiment reproducibility possible without bloating the Git repository
- Version numbers in filenames are an anti-pattern — use Git for version control and keep one canonical filename for each artifact