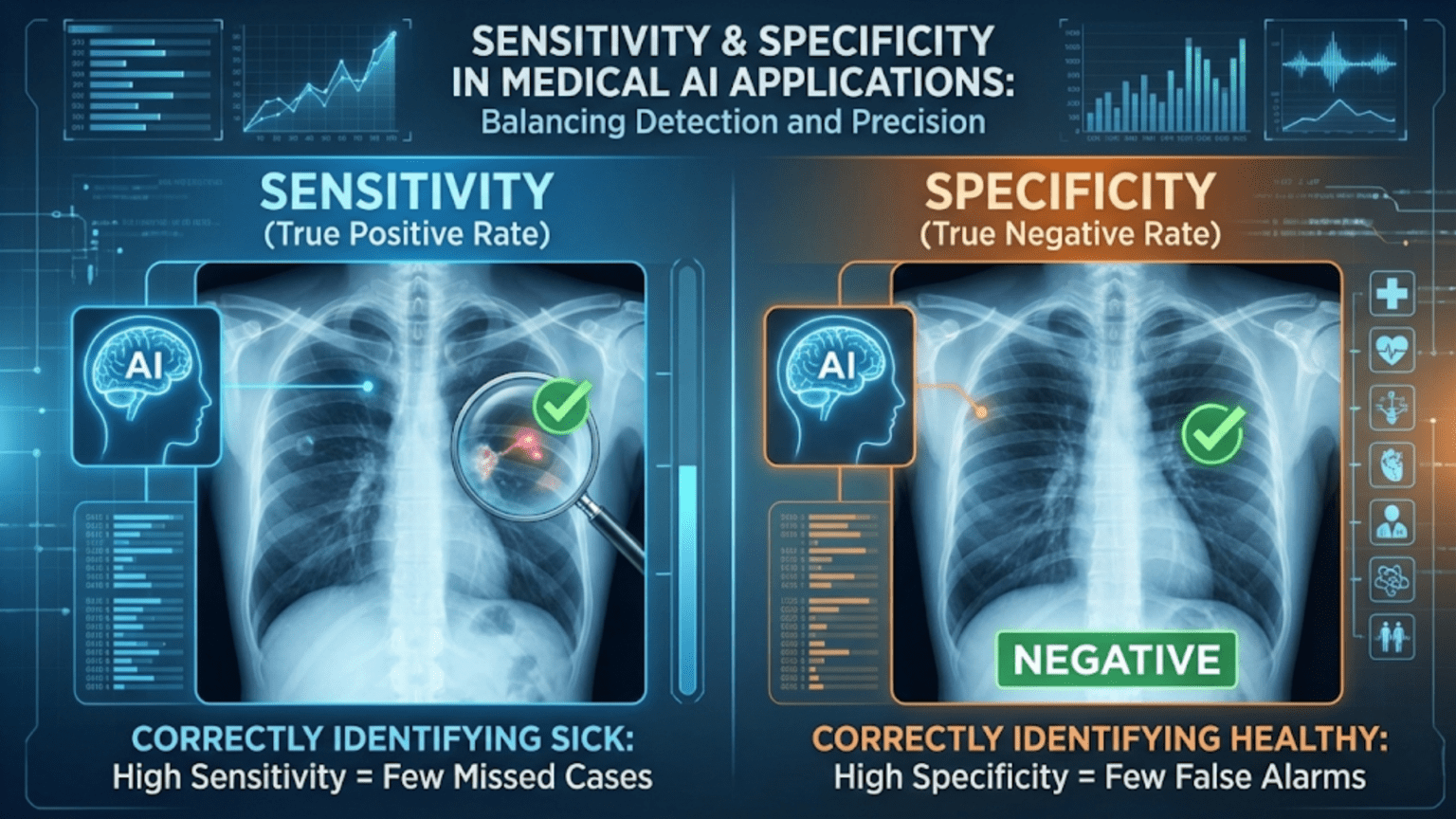
Sensitivity (also called recall or the True Positive Rate) measures how well a medical AI correctly identifies patients who have a condition — it answers “of all sick patients, how many did the test correctly flag?” Specificity (the True Negative Rate) measures how well it correctly identifies healthy patients — it answers “of all healthy patients, how many did the test correctly clear?” Together, they are the two most fundamental metrics in clinical diagnostics and medical AI evaluation, and understanding the tradeoff between them is essential for safe deployment of AI in healthcare.
Introduction
In 2020, a deep learning model for detecting diabetic retinopathy was deployed in a clinic in Thailand. The model had high sensitivity — it caught the vast majority of patients with serious eye damage. But it also had a high rate of false positives, generating anxiety among patients who were later found to be healthy. Clinicians had to recalibrate how they presented the model’s findings, adjusting the decision threshold and modifying their communication protocols.
This scenario plays out in medical AI deployments around the world. A model that performs brilliantly on a benchmark dataset can have very different real-world consequences depending on how its sensitivity and specificity are calibrated — and those two metrics, not accuracy alone, determine whether a medical AI helps or harms patients.
Sensitivity and specificity are the language of medical diagnostics. They are how clinicians, regulators, and ethicists evaluate diagnostic tests — and they are therefore how medical AI systems must be evaluated. Unlike generic machine learning metrics, they carry specific clinical interpretations that connect model behavior directly to patient outcomes.
This article builds a complete understanding of sensitivity and specificity in the medical context: their definitions, the clinical meaning of their tradeoffs, how prevalence affects what a test result means (Bayes’ theorem in action), regulatory requirements, and a complete Python toolkit for medical AI evaluation.
From Confusion Matrix to Clinical Language
Medical AI uses the same underlying four-outcome framework as all binary classification, but with domain-specific terminology that reflects clinical context.
In medical testing:
- Positive = disease or condition is present
- Negative = disease or condition is absent
The four outcomes map directly to clinical events:
| General Term | Medical Term | Clinical Meaning |
|---|---|---|
| True Positive (TP) | True Positive | Test correctly identifies a sick patient |
| True Negative (TN) | True Negative | Test correctly clears a healthy patient |
| False Positive (FP) | False Positive | Test incorrectly flags a healthy patient as sick |
| False Negative (FN) | False Negative | Test incorrectly clears a sick patient as healthy |
From these four counts, sensitivity and specificity are derived directly:
Sensitivity: Never Miss a Sick Patient
Definition and Interpretation
Sensitivity is the probability that the test correctly identifies a patient who truly has the condition. It answers the question: “If a patient is sick, how likely is the test to detect it?”
A sensitivity of 0.95 (95%) means that for every 100 patients who actually have the disease, the test correctly flags 95 and misses 5. Those 5 missed patients (false negatives) leave the clinic believing they are healthy when they are not.
Sensitivity is identical to recall and the True Positive Rate (TPR) in machine learning terminology. The different names reflect different communities using the same concept:
- Clinicians: sensitivity
- Epidemiologists: true positive rate
- Machine learning researchers: recall
- Signal detection theorists: hit rate
All refer to the same calculation: TP / (TP + FN).
When Sensitivity Is the Primary Concern
Sensitivity is paramount when the consequences of missing a real case are severe. In clinical settings, this typically means:
Life-threatening conditions: Cancer, HIV, tuberculosis, sepsis. Missing an early diagnosis of cancer can allow it to progress to an untreatable stage. A highly sensitive screening test catches nearly all cases, even at the cost of some false alarms that will be resolved by follow-up testing.
Infectious diseases: Missing a case of a contagious disease means an infected patient goes untreated and unknowingly exposes others. During epidemic response, maximizing sensitivity of screening tools is a public health imperative.
Emergency triage: When a patient presents in the emergency department with symptoms that could indicate a cardiac event, high sensitivity is critical. It is far better to admit patients who turn out not to have had a heart attack than to send home a patient who is actually experiencing one.
Blood supply safety: Testing donated blood for HIV, hepatitis B and C, and other pathogens requires extremely high sensitivity. A false negative could contaminate the blood supply and harm recipients.
Specificity: Never Alarm a Healthy Patient
Definition and Interpretation
Specificity is the probability that the test correctly clears a patient who is truly healthy. It answers: “If a patient is healthy, how likely is the test to confirm that?”
A specificity of 0.90 (90%) means that for every 100 healthy patients tested, 90 are correctly cleared and 10 are incorrectly flagged as potentially sick. Those 10 false positives will face unnecessary follow-up tests, anxiety, potential side effects from additional procedures, and healthcare costs.
Specificity is identical to the True Negative Rate (TNR) and equal to (1 − False Positive Rate).
When Specificity Is the Primary Concern
Specificity is paramount when the consequences of false alarms are severe. In clinical settings:
Confirmatory testing: Before an irreversible or high-risk intervention (surgery, chemotherapy, lifelong medication), you need a highly specific test to confirm the diagnosis. You cannot afford to put healthy patients through aggressive treatment.
Rare disease screening: When prevalence is very low, even a modest false positive rate generates many more false alarms than true detections (we will quantify this precisely using Bayes’ theorem below). High specificity prevents overwhelming the healthcare system with unnecessary follow-up.
Stigmatizing diagnoses: Some diagnoses carry significant social and psychological consequences — HIV, certain mental health conditions, genetic predispositions. A false positive can cause lasting harm to a healthy person.
High-cost follow-up procedures: If a positive screening test leads to expensive, uncomfortable, or risky follow-up procedures (colonoscopy, lumbar puncture, cardiac catheterization), unnecessary false positives impose meaningful costs and risks on healthy individuals.
The Sensitivity-Specificity Tradeoff
Like precision and recall, sensitivity and specificity exist in a tradeoff. For any classifier that outputs probability scores, increasing sensitivity always comes at the cost of specificity, and vice versa. Adjusting the decision threshold is how this tradeoff is navigated.
Visualizing the Tradeoff
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve, roc_auc_score
# Simulate a medical AI diagnostic dataset
# 1000 patients, 20% disease prevalence (200 sick, 800 healthy)
np.random.seed(42)
X, y = make_classification(
n_samples=2000,
n_features=15,
n_informative=10,
weights=[0.80, 0.20], # 80% healthy, 20% diseased
random_state=42
)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=42, stratify=y
)
# Train logistic regression (simulating a medical AI model)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_s = scaler.fit_transform(X_train)
X_test_s = scaler.transform(X_test)
model = LogisticRegression(random_state=42, max_iter=1000, class_weight='balanced')
model.fit(X_train_s, y_train)
y_proba = model.predict_proba(X_test_s)[:, 1]
# Compute sensitivity and specificity at each threshold
fpr, tpr, thresholds = roc_curve(y_test, y_proba)
sensitivity = tpr # TPR = sensitivity
specificity = 1 - fpr # TNR = specificity = 1 - FPR
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
# Panel 1: Sensitivity and Specificity vs Threshold
axes[0].plot(thresholds, sensitivity[:-1], 'b-', linewidth=2.5, label='Sensitivity (TPR)')
axes[0].plot(thresholds, specificity[:-1], 'r-', linewidth=2.5, label='Specificity (TNR)')
axes[0].axvline(x=0.5, color='gray', linestyle='--', alpha=0.7, label='Default threshold (0.5)')
# Find where sensitivity ≈ specificity (crossover point)
diff = np.abs(sensitivity[:-1] - specificity[:-1])
crossover_idx = np.argmin(diff)
crossover_t = thresholds[crossover_idx]
axes[0].axvline(x=crossover_t, color='green', linestyle=':', linewidth=2,
label=f'Crossover threshold ({crossover_t:.3f})')
axes[0].set_xlabel("Decision Threshold", fontsize=12)
axes[0].set_ylabel("Rate", fontsize=12)
axes[0].set_title("Sensitivity & Specificity vs Threshold\n"
"(Moving threshold right ↑ specificity ↓ sensitivity)",
fontsize=12, fontweight='bold')
axes[0].legend(fontsize=10)
axes[0].grid(True, alpha=0.3)
axes[0].set_xlim([0, 1])
axes[0].set_ylim([0, 1.05])
# Panel 2: ROC Curve (= Sensitivity vs 1-Specificity)
auc_score = roc_auc_score(y_test, y_proba)
axes[1].plot(fpr, tpr, 'b-', linewidth=2.5, label=f'ROC Curve (AUC={auc_score:.3f})')
axes[1].plot([0, 1], [0, 1], 'k--', linewidth=1.5, label='Random classifier')
axes[1].scatter(1 - specificity[crossover_idx], sensitivity[crossover_idx],
color='green', s=200, zorder=5,
label=f'Crossover point\n(Sens=Spec≈{sensitivity[crossover_idx]:.3f})')
axes[1].set_xlabel("1 - Specificity (False Positive Rate)", fontsize=12)
axes[1].set_ylabel("Sensitivity (True Positive Rate)", fontsize=12)
axes[1].set_title("ROC Curve = Sensitivity vs (1 - Specificity)\n"
"(AUC summarizes the tradeoff across all thresholds)",
fontsize=12, fontweight='bold')
axes[1].legend(fontsize=10, loc='lower right')
axes[1].grid(True, alpha=0.3)
plt.suptitle("The Sensitivity-Specificity Tradeoff", fontsize=14, fontweight='bold', y=1.01)
plt.tight_layout()
plt.savefig("sensitivity_specificity_tradeoff.png", dpi=150, bbox_inches='tight')
plt.show()
print("Saved: sensitivity_specificity_tradeoff.png")
# Print values at clinically relevant thresholds
print(f"\n{'Threshold':>10} | {'Sensitivity':>12} | {'Specificity':>12} | {'Sum':>6}")
print("-" * 50)
for t in [0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8]:
idx = np.argmin(np.abs(thresholds - t))
sens = sensitivity[idx]
spec = specificity[idx]
print(f"{t:>10.1f} | {sens:>12.4f} | {spec:>12.4f} | {sens+spec:>6.4f}")The ROC curve is the visual representation of the complete sensitivity-specificity tradeoff. Every point on the curve corresponds to a specific threshold, with its y-coordinate being sensitivity and its x-coordinate being (1 − specificity). The AUC summarizes model quality across all possible operating points.
Bayes’ Theorem: Why Prevalence Changes Everything
This is the concept most frequently misunderstood by beginners — and one of the most practically important ideas in all of medical AI.
Even a highly sensitive and specific test will produce a large proportion of false positives when the disease prevalence is very low. This is not a flaw in the test; it is a mathematical inevitability. Understanding this prevents costly misinterpretations of medical test results.
Positive Predictive Value (PPV) and Negative Predictive Value (NPV)
Two additional metrics complete the picture:
Positive Predictive Value (PPV): Given that the test says positive, what is the probability the patient actually has the disease?
Negative Predictive Value (NPV): Given that the test says negative, what is the probability the patient is actually healthy?
Unlike sensitivity and specificity, PPV and NPV depend critically on disease prevalence. A test with fixed sensitivity and specificity will have very different PPV values when applied to populations with different disease rates.
The Impact of Prevalence: A Worked Example
Consider a COVID-19 screening test with:
- Sensitivity = 90% (catches 90% of infected people)
- Specificity = 95% (correctly clears 95% of healthy people)
These sound excellent. Let’s calculate PPV under three different prevalence scenarios:
import numpy as np
import matplotlib.pyplot as plt
def calculate_ppv_npv(sensitivity, specificity, prevalence):
"""
Calculate PPV, NPV, and confusion matrix for a given test
and disease prevalence using Bayes' theorem.
Args:
sensitivity: True Positive Rate (TP / (TP + FN))
specificity: True Negative Rate (TN / (TN + FP))
prevalence: Probability of disease in the tested population
Returns:
Dictionary of all metrics
"""
# For a population of 10,000 people
n = 10000
n_diseased = int(n * prevalence)
n_healthy = n - n_diseased
TP = int(n_diseased * sensitivity)
FN = n_diseased - TP
TN = int(n_healthy * specificity)
FP = n_healthy - TN
# PPV = P(disease | test positive) = TP / (TP + FP)
ppv = TP / (TP + FP) if (TP + FP) > 0 else 0
# NPV = P(no disease | test negative) = TN / (TN + FN)
npv = TN / (TN + FN) if (TN + FN) > 0 else 0
# Alternatively via Bayes' theorem (same result):
# PPV = (sensitivity * prevalence) / (sensitivity * prevalence + (1-specificity) * (1-prevalence))
ppv_bayes = (sensitivity * prevalence) / (
sensitivity * prevalence + (1 - specificity) * (1 - prevalence)
)
return {
"prevalence": prevalence,
"n_diseased": n_diseased,
"n_healthy": n_healthy,
"TP": TP,
"FP": FP,
"FN": FN,
"TN": TN,
"PPV": ppv,
"NPV": npv,
"PPV_bayes": ppv_bayes,
"false_alarm_ratio": FP / (TP + FP) if (TP + FP) > 0 else 0
}
# Test: sensitivity=90%, specificity=95%
sensitivity = 0.90
specificity = 0.95
print("=== COVID-19 Test (Sensitivity=90%, Specificity=95%) ===")
print(" Population size: 10,000 people per scenario\n")
print(f"{'Prevalence':>12} | {'Diseased':>9} | {'TP':>6} | {'FP':>6} | {'FN':>6} | {'PPV':>8} | {'NPV':>8} | {'Of pos results, % false'}")
print("-" * 100)
for prev in [0.001, 0.01, 0.05, 0.10, 0.20, 0.50]:
r = calculate_ppv_npv(sensitivity, specificity, prev)
print(f"{prev*100:>11.1f}% | {r['n_diseased']:>9,} | {r['TP']:>6} | {r['FP']:>6} | "
f"{r['FN']:>6} | {r['PPV']:>8.4f} | {r['NPV']:>8.4f} | {r['false_alarm_ratio']*100:>22.1f}%")The results reveal a striking and counterintuitive pattern. At 0.1% prevalence (a general population screen), even with a 95%-specific test, 83% of positive results are false alarms. At 1% prevalence, still more than half of positive results are false. Only when prevalence reaches 10–20% (a high-risk population) does the PPV become acceptably high.
This is why screening programs are designed in stages: first screen a broad population with a highly sensitive test to identify candidates, then confirm positives with a highly specific test. The sensitivity of the first stage maximizes the chance of catching all true cases; the specificity of the second stage minimizes unnecessary treatment.
Visualizing How PPV Changes with Prevalence
import numpy as np
import matplotlib.pyplot as plt
prevalences = np.linspace(0.001, 0.5, 500)
# Several test quality levels
test_configs = [
(0.90, 0.90, "Sens=90%, Spec=90%"),
(0.90, 0.95, "Sens=90%, Spec=95%"),
(0.95, 0.95, "Sens=95%, Spec=95%"),
(0.99, 0.99, "Sens=99%, Spec=99%"),
]
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
# Panel 1: PPV vs Prevalence
for sens, spec, label in test_configs:
ppv_vals = [(sens * p) / (sens * p + (1 - spec) * (1 - p)) for p in prevalences]
axes[0].plot(prevalences * 100, ppv_vals, linewidth=2.5, label=label)
axes[0].axhline(y=0.5, color='red', linestyle='--', alpha=0.5, label='PPV = 0.5 line')
axes[0].set_xlabel("Disease Prevalence (%)", fontsize=12)
axes[0].set_ylabel("Positive Predictive Value (PPV)", fontsize=12)
axes[0].set_title("PPV vs Prevalence\n(Even excellent tests have low PPV at low prevalence)",
fontsize=12, fontweight='bold')
axes[0].legend(fontsize=9)
axes[0].grid(True, alpha=0.3)
axes[0].set_ylim([0, 1.05])
# Panel 2: NPV vs Prevalence
for sens, spec, label in test_configs:
npv_vals = [(spec * (1-p)) / (spec * (1-p) + (1-sens) * p) for p in prevalences]
axes[1].plot(prevalences * 100, npv_vals, linewidth=2.5, label=label)
axes[1].set_xlabel("Disease Prevalence (%)", fontsize=12)
axes[1].set_ylabel("Negative Predictive Value (NPV)", fontsize=12)
axes[1].set_title("NPV vs Prevalence\n(NPV drops as prevalence rises — more false reassurances)",
fontsize=12, fontweight='bold')
axes[1].legend(fontsize=9)
axes[1].grid(True, alpha=0.3)
axes[1].set_ylim([0, 1.05])
plt.suptitle("How Disease Prevalence Affects What Test Results Mean",
fontsize=14, fontweight='bold', y=1.01)
plt.tight_layout()
plt.savefig("ppv_npv_vs_prevalence.png", dpi=150, bbox_inches='tight')
plt.show()
print("Saved: ppv_npv_vs_prevalence.png")Likelihood Ratios: A Powerful Clinical Tool
A limitation of PPV and NPV is that they change with every population. A more portable measure is the Likelihood Ratio (LR), which quantifies how much a test result updates your probability estimate of disease.
Positive Likelihood Ratio (LR+)
How much more likely is a positive test result in a sick patient than in a healthy patient?
A LR+ of 10 means a positive test result is 10 times more likely in a diseased patient than in a healthy one. Values above 10 provide strong evidence for disease.
Negative Likelihood Ratio (LR−)
How much more likely is a negative test result in a healthy patient than in a sick one?
A LR− of 0.1 means a negative test result is 10 times more likely in a healthy patient. Values below 0.1 provide strong evidence against disease.
import numpy as np
def likelihood_ratios(sensitivity, specificity):
"""Compute positive and negative likelihood ratios."""
lr_pos = sensitivity / (1 - specificity) if (1 - specificity) > 0 else float('inf')
lr_neg = (1 - sensitivity) / specificity if specificity > 0 else 0
return lr_pos, lr_neg
def interpret_lr(lr):
"""Provide clinical interpretation of a likelihood ratio."""
if lr >= 10:
return "Strong evidence FOR disease"
elif lr >= 5:
return "Moderate evidence FOR disease"
elif lr >= 2:
return "Weak evidence for disease"
elif lr > 0.5:
return "Minimal evidence (limited clinical value)"
elif lr > 0.2:
return "Weak evidence AGAINST disease"
elif lr > 0.1:
return "Moderate evidence AGAINST disease"
else:
return "Strong evidence AGAINST disease"
# Compare several medical AI models
test_scenarios = [
("Mammography (screening)", 0.75, 0.92),
("PSA Test (prostate cancer)", 0.80, 0.70),
("CT Pulmonary Angiography", 0.83, 0.96),
("AI Skin Lesion Classifier", 0.90, 0.85),
("AI Diabetic Retinopathy", 0.97, 0.87),
("AI Chest X-ray (COVID)", 0.88, 0.91),
("PCR Test (near-perfect)", 0.99, 0.99),
]
print("=== Likelihood Ratios for Medical AI Systems ===\n")
print(f"{'Test / Model':<35} | {'Sens':>6} | {'Spec':>6} | {'LR+':>7} | {'LR-':>7} | {'LR+ Interpretation'}")
print("-" * 100)
for name, sens, spec in test_scenarios:
lr_p, lr_n = likelihood_ratios(sens, spec)
interp = interpret_lr(lr_p)
print(f"{name:<35} | {sens:>6.2f} | {spec:>6.2f} | {lr_p:>7.2f} | {lr_n:>7.4f} | {interp}")
print("\nRule of thumb:")
print(" LR+ > 10 → Strong evidence of disease when test is positive")
print(" LR- < 0.1 → Strong evidence against disease when test is negative")Medical AI in Practice: Key Application Areas
1. Radiology AI: Detecting Disease in Images
Radiology represents one of the most advanced and clinically validated uses of AI. Models reading chest X-rays, mammograms, CT scans, and MRIs must balance sensitivity and specificity against workflow constraints.
Example: Chest X-ray AI for Pneumonia
A radiologist reviews hundreds of chest X-rays per day. An AI tool designed to flag urgent cases needs extremely high sensitivity (don’t miss pneumonia, especially in critically ill patients) but can tolerate moderate false positive rates because the radiologist will perform a final review.
For a routine screening tool (annual chest X-ray), higher specificity is needed to avoid overwhelming radiologists with false positives on a large, mostly-healthy population.
from sklearn.metrics import confusion_matrix, roc_curve, roc_auc_score
import numpy as np
import matplotlib.pyplot as plt
def medical_ai_evaluation(y_true, y_proba, model_name,
condition_name="Disease",
high_sensitivity_threshold=0.20,
balanced_threshold=0.50,
high_specificity_threshold=0.80):
"""
Complete medical AI evaluation with clinical context.
Evaluates model at three clinically relevant thresholds:
1. High-sensitivity: maximizes disease detection (emergency/triage)
2. Balanced: default operating point
3. High-specificity: minimizes false alarms (confirmatory/rare disease)
"""
from sklearn.metrics import roc_auc_score
auc = roc_auc_score(y_true, y_proba)
print(f"\n{'='*60}")
print(f" Medical AI Evaluation: {model_name}")
print(f" Condition: {condition_name}")
print(f" AUC-ROC: {auc:.4f}")
print(f"{'='*60}")
n_pos = y_true.sum()
n_neg = len(y_true) - n_pos
prevalence = n_pos / len(y_true)
print(f"\n Test population: {len(y_true):,} patients")
print(f" Diseased: {n_pos:,} ({prevalence*100:.1f}%)")
print(f" Healthy: {n_neg:,} ({(1-prevalence)*100:.1f}%)")
print(f"\n {'Threshold':>12} | {'Sensitivity':>12} | {'Specificity':>12} | "
f"{'PPV':>6} | {'NPV':>6} | {'Use Case'}")
print("-" * 90)
for threshold, use_case in [
(high_sensitivity_threshold, "Emergency / Triage (miss nothing)"),
(balanced_threshold, "General screening"),
(high_specificity_threshold, "Confirmatory (minimize false alarms)"),
]:
y_pred = (y_proba >= threshold).astype(int)
cm = confusion_matrix(y_true, y_pred)
# Handle case where all predictions are same class
if cm.shape == (1, 2) or cm.shape == (2, 1):
continue
tn, fp, fn, tp = cm.ravel()
sens = tp / (tp + fn) if (tp + fn) > 0 else 0
spec = tn / (tn + fp) if (tn + fp) > 0 else 0
ppv = tp / (tp + fp) if (tp + fp) > 0 else 0
npv = tn / (tn + fn) if (tn + fn) > 0 else 0
print(f" {threshold:>12.2f} | {sens:>12.4f} | {spec:>12.4f} | "
f"{ppv:>6.4f} | {npv:>6.4f} | {use_case}")
return auc
# Simulate chest X-ray pneumonia detection model
np.random.seed(42)
X_cxr, y_cxr = make_classification(
n_samples=5000,
n_features=20,
n_informative=12,
weights=[0.85, 0.15], # 15% pneumonia prevalence
random_state=42
)
X_tr_cxr, X_te_cxr, y_tr_cxr, y_te_cxr = train_test_split(
X_cxr, y_cxr, test_size=0.25, random_state=42, stratify=y_cxr
)
scaler_cxr = StandardScaler()
X_tr_s_cxr = scaler_cxr.fit_transform(X_tr_cxr)
X_te_s_cxr = scaler_cxr.transform(X_te_cxr)
from sklearn.ensemble import GradientBoostingClassifier
cxr_model = GradientBoostingClassifier(n_estimators=200, random_state=42)
cxr_model.fit(X_tr_s_cxr, y_tr_cxr)
y_proba_cxr = cxr_model.predict_proba(X_te_s_cxr)[:, 1]
medical_ai_evaluation(y_te_cxr, y_proba_cxr,
model_name="Chest X-ray Pneumonia Detector",
condition_name="Pneumonia")2. Cancer Screening: The Two-Stage Paradigm
Most cancer screening programs use a two-stage approach that leverages sensitivity and specificity optimally at each stage:
Stage 1 (Broad Screening): Apply a highly sensitive test to a large population. The goal is to catch virtually all true cases. False positives at this stage are acceptable because they will be caught by the confirmatory test.
Stage 2 (Confirmatory Testing): Apply a highly specific test only to Stage 1 positives. The goal is to confirm true cases while clearing false positives from Stage 1.
import numpy as np
def two_stage_screening_simulation(population_size, prevalence,
stage1_sensitivity, stage1_specificity,
stage2_sensitivity, stage2_specificity,
condition="Cancer"):
"""
Simulate a two-stage screening program and report outcomes.
Args:
population_size: Total population screened
prevalence: True disease prevalence
stage1_sensitivity: Sensitivity of first screening test
stage1_specificity: Specificity of first screening test
stage2_sensitivity: Sensitivity of confirmatory test
stage2_specificity: Specificity of confirmatory test
condition: Name of the condition being screened
"""
n_diseased = int(population_size * prevalence)
n_healthy = population_size - n_diseased
# --- Stage 1: Broad Screening ---
s1_TP = int(n_diseased * stage1_sensitivity) # Diseased flagged
s1_FN = n_diseased - s1_TP # Diseased missed (exit screening)
s1_FP = int(n_healthy * (1 - stage1_specificity)) # Healthy flagged
s1_TN = n_healthy - s1_FP # Healthy cleared (exit screening)
# Those who proceed to Stage 2 = Stage 1 positives
s2_population_diseased = s1_TP # True positives from Stage 1
s2_population_healthy = s1_FP # False positives from Stage 1
# --- Stage 2: Confirmatory Testing ---
s2_TP = int(s2_population_diseased * stage2_sensitivity)
s2_FN = s2_population_diseased - s2_TP # Missed by Stage 2 (false clear)
s2_TN = int(s2_population_healthy * stage2_specificity)
s2_FP = s2_population_healthy - s2_TN # Still flagged after Stage 2
# Final outcomes across the full program
final_TP = s2_TP # Correctly diagnosed (receive treatment)
final_FN = s1_FN + s2_FN # Total missed cases
final_FP = s2_FP # Healthy patients who complete both stages
final_TN = s1_TN + s2_TN # Correctly cleared at either stage
overall_sensitivity = final_TP / n_diseased if n_diseased > 0 else 0
overall_specificity = final_TN / n_healthy if n_healthy > 0 else 0
overall_ppv = final_TP / (final_TP + final_FP) if (final_TP + final_FP) > 0 else 0
print(f"\n{'='*60}")
print(f" Two-Stage {condition} Screening Program")
print(f"{'='*60}")
print(f" Population: {population_size:,} people")
print(f" True prevalence: {prevalence*100:.2f}% ({n_diseased:,} cases)")
print(f"\n Stage 1 (Broad Screen: Sens={stage1_sensitivity:.0%}, Spec={stage1_specificity:.0%}):")
print(f" {s1_TP + s1_FP:,} flagged for Stage 2 ({s1_TP:,} true + {s1_FP:,} false)")
print(f" {s1_FN:,} diseased patients missed at Stage 1 ← serious")
print(f" {s1_TN:,} healthy patients correctly cleared")
print(f"\n Stage 2 (Confirmatory: Sens={stage2_sensitivity:.0%}, Spec={stage2_specificity:.0%}):")
print(f" Of {s1_TP + s1_FP:,} Stage 1 positives tested:")
print(f" {s2_TP:,} confirmed diseased (receive treatment)")
print(f" {s2_TN:,} healthy patients cleared at Stage 2")
print(f" {s2_FP:,} healthy patients still falsely confirmed ← proceed to treatment")
print(f" {s2_FN:,} additional diseased patients missed at Stage 2")
print(f"\n Overall Program Performance:")
print(f" Overall sensitivity: {overall_sensitivity:.4f} ({final_FN:,} diseased patients missed)")
print(f" Overall specificity: {overall_specificity:.4f} ({final_FP:,} healthy patients treated unnecessarily)")
print(f" Overall PPV: {overall_ppv:.4f} ({final_TP/(final_TP+final_FP)*100:.1f}% of treated patients truly diseased)")
print(f"\n ✓ Treated correctly: {final_TP:,}")
print(f" ✗ Missed cases: {final_FN:,}")
print(f" ✗ Over-treated: {final_FP:,}")
# Breast cancer screening program
two_stage_screening_simulation(
population_size = 100_000,
prevalence = 0.01, # 1% prevalence (1,000 cases)
stage1_sensitivity = 0.90, # Mammography sensitivity
stage1_specificity = 0.85, # Mammography specificity
stage2_sensitivity = 0.95, # Biopsy sensitivity
stage2_specificity = 0.99, # Biopsy specificity
condition = "Breast Cancer"
)Regulatory Context: What Authorities Require
Medical AI systems are regulated as medical devices in most jurisdictions. Understanding what regulators require around sensitivity and specificity is essential for anyone building clinical AI.
FDA (United States)
The FDA regulates AI-based medical devices under the Software as a Medical Device (SaMD) framework. Key requirements include:
- Pre-defined performance targets: Sponsors must specify minimum acceptable sensitivity and specificity before validation studies begin.
- Confidence intervals: Point estimates (e.g., “sensitivity = 92%”) are insufficient. Studies must report 95% confidence intervals.
- Subgroup analysis: Performance must be validated across relevant demographic subgroups (age, sex, race/ethnicity) to detect disparities.
- Intended use alignment: Performance standards are calibrated to the intended use case (screening vs. confirmatory testing).
CE Marking (European Union)
Under the Medical Device Regulation (MDR), AI diagnostic tools must demonstrate:
- Clinical performance validation with sensitivity/specificity benchmarked against predicate devices
- Performance in the intended population and care setting
- Monitoring requirements for real-world performance drift
Interpreting Performance Claims
When a medical AI company claims “95% sensitivity and 93% specificity,” several questions must be answered:
- What population was this measured in? (Hospital referrals vs. general screening populations have very different disease prevalences)
- What was the comparator (gold standard)? (Expert consensus, biopsy, PCR test?)
- What confidence intervals were reported?
- Were results independently validated?
- How does performance compare in the specific demographic groups relevant to your setting?
import numpy as np
from scipy import stats
def compute_performance_with_ci(TP, FP, FN, TN, confidence=0.95):
"""
Compute sensitivity, specificity, PPV, NPV and their
exact binomial confidence intervals (Wilson method).
Args:
TP, FP, FN, TN: Confusion matrix values
confidence: Confidence level (default 0.95 = 95% CI)
Returns:
DataFrame-like dictionary of metrics with confidence intervals
"""
from statsmodels.stats.proportion import proportion_confint
z = stats.norm.ppf((1 + confidence) / 2)
def wilson_ci(k, n, z):
"""Wilson score confidence interval for a proportion."""
if n == 0:
return 0.0, 0.0
p = k / n
center = (p + z**2 / (2*n)) / (1 + z**2 / n)
margin = (z * np.sqrt(p*(1-p)/n + z**2/(4*n**2))) / (1 + z**2/n)
return max(0, center - margin), min(1, center + margin)
n_pos = TP + FN # Total actual positives
n_neg = FP + TN # Total actual negatives
n_pred_pos = TP + FP
n_pred_neg = FN + TN
metrics = {}
for name, k, n in [
("Sensitivity (Recall)", TP, n_pos),
("Specificity", TN, n_neg),
("PPV (Precision)", TP, n_pred_pos),
("NPV", TN, n_pred_neg),
]:
point_est = k / n if n > 0 else 0
lo, hi = wilson_ci(k, n, z)
metrics[name] = {
"value": point_est,
"lower": lo,
"upper": hi,
"n": n
}
return metrics
def print_clinical_report(TP, FP, FN, TN, model_name="Medical AI Model",
condition="Disease", confidence=0.95):
"""
Print a clinical-style performance report with confidence intervals.
This format mirrors what would appear in a medical AI validation study.
"""
print(f"\n{'='*65}")
print(f" CLINICAL PERFORMANCE REPORT")
print(f" Model: {model_name}")
print(f" Condition: {condition}")
print(f"{'='*65}")
n_total = TP + FP + FN + TN
n_pos = TP + FN
n_neg = FP + TN
prevalence = n_pos / n_total if n_total > 0 else 0
print(f"\n Study Population: {n_total:,} patients")
print(f" Disease-positive: {n_pos:,} ({prevalence*100:.1f}%)")
print(f" Disease-negative: {n_neg:,} ({(1-prevalence)*100:.1f}%)")
print(f"\n Confusion Matrix:")
print(f" {'':25} Predicted Positive Predicted Negative")
print(f" {'Actual Positive':<25} {TP:>18,} {FN:>18,}")
print(f" {'Actual Negative':<25} {FP:>18,} {TN:>18,}")
metrics = compute_performance_with_ci(TP, FP, FN, TN, confidence)
ci_pct = int(confidence * 100)
print(f"\n Performance Metrics ({ci_pct}% Confidence Intervals):\n")
print(f" {'Metric':<28} | {'Value':>8} | {ci_pct}% CI")
print("-" * 55)
for name, m in metrics.items():
print(f" {name:<28} | {m['value']:>8.4f} | "
f"[{m['lower']:.4f}, {m['upper']:.4f}] (n={m['n']:,})")
# Likelihood ratios
sens = metrics["Sensitivity (Recall)"]["value"]
spec = metrics["Specificity"]["value"]
fpr = 1 - spec
lr_pos = sens / fpr if fpr > 0 else float('inf')
lr_neg = (1 - sens) / spec if spec > 0 else 0
print(f"\n Likelihood Ratios:")
print(f" LR+ = {lr_pos:.2f} (positive test increases disease odds by {lr_pos:.1f}×)")
print(f" LR- = {lr_neg:.4f} (negative test reduces disease odds to {lr_neg*100:.1f}% of prior)")
# Simulate a 2,000-patient validation study for a skin cancer AI
print_clinical_report(
TP=342, FP=89, FN=28, TN=1541,
model_name="DermAI v2.0",
condition="Malignant Melanoma"
)Subgroup Analysis: Fairness in Medical AI
A critical and often overlooked aspect of medical AI evaluation is performance across demographic subgroups. A model with 92% sensitivity overall may have 98% sensitivity in one demographic group and only 78% in another — a disparity that could constitute a dangerous health equity failure.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import confusion_matrix
def subgroup_performance_analysis(model, X_test, y_test, groups,
group_names, scaler=None, model_name="Model"):
"""
Analyze model performance across demographic subgroups.
Reports sensitivity, specificity, and PPV for each group.
Args:
model: Trained classifier
X_test: Test features
y_test: True labels
groups: Array of group assignments for each test sample
group_names: Names corresponding to group IDs
scaler: Optional scaler for feature preprocessing
model_name: Name for display
"""
X_scaled = scaler.transform(X_test) if scaler else X_test
y_proba = model.predict_proba(X_scaled)[:, 1]
y_pred = (y_proba >= 0.5).astype(int)
print(f"\n=== Subgroup Analysis: {model_name} ===\n")
print(f"{'Subgroup':<20} | {'N':>6} | {'Prevalence':>11} | {'Sensitivity':>12} | "
f"{'Specificity':>12} | {'PPV':>8}")
print("-" * 80)
# Overall
cm_all = confusion_matrix(y_test, y_pred)
tn_a, fp_a, fn_a, tp_a = cm_all.ravel()
sens_a = tp_a / (tp_a + fn_a)
spec_a = tn_a / (tn_a + fp_a)
ppv_a = tp_a / (tp_a + fp_a) if (tp_a + fp_a) > 0 else 0
prev_a = (tp_a + fn_a) / len(y_test)
print(f"{'OVERALL':<20} | {len(y_test):>6,} | {prev_a:>10.1%} | {sens_a:>12.4f} | "
f"{spec_a:>12.4f} | {ppv_a:>8.4f}")
print("-" * 80)
disparities = []
for gid, gname in enumerate(group_names):
mask = (groups == gid)
if mask.sum() < 10:
continue
X_g = X_test[mask]
y_g = y_test[mask]
y_pred_g = y_pred[mask]
cm_g = confusion_matrix(y_g, y_pred_g, labels=[0, 1])
if cm_g.shape != (2, 2):
continue
tn_g, fp_g, fn_g, tp_g = cm_g.ravel()
sens_g = tp_g / (tp_g + fn_g) if (tp_g + fn_g) > 0 else 0
spec_g = tn_g / (tn_g + fp_g) if (tn_g + fp_g) > 0 else 0
ppv_g = tp_g / (tp_g + fp_g) if (tp_g + fp_g) > 0 else 0
prev_g = (tp_g + fn_g) / len(y_g) if len(y_g) > 0 else 0
flag = "⚠" if abs(sens_g - sens_a) > 0.05 else " "
print(f"{gname:<20} | {mask.sum():>6,} | {prev_g:>10.1%} | "
f"{sens_g:>12.4f} | {spec_g:>12.4f} | {ppv_g:>8.4f} {flag}")
disparities.append((gname, sens_g, spec_g, ppv_g))
# Check for significant disparities
if disparities:
sens_vals = [d[1] for d in disparities]
max_gap = max(sens_vals) - min(sens_vals)
print(f"\n Sensitivity range across groups: {min(sens_vals):.4f} – {max(sens_vals):.4f}")
print(f" Max sensitivity gap: {max_gap:.4f}", end="")
if max_gap > 0.05:
print(f" ⚠ DISPARITY DETECTED (>{5}% gap)")
else:
print(f" ✓ No significant disparity")
print("\n ⚠ = sensitivity gap > 5% from overall average")
# Simulate demographic disparities
np.random.seed(42)
n = 3000
# Generate base features
X_base, y_base = make_classification(
n_samples=n, n_features=15, n_informative=10,
weights=[0.75, 0.25], random_state=42
)
# Simulate group membership (3 groups: 40%, 35%, 25%)
groups = np.random.choice([0, 1, 2], size=n, p=[0.40, 0.35, 0.25])
# Add group-specific feature noise (simulating real-world disparities)
for g_idx, noise_level in [(1, 0.3), (2, 0.8)]:
mask = (groups == g_idx)
X_base[mask] += np.random.normal(0, noise_level, (mask.sum(), X_base.shape[1]))
group_names = ["Group A (40%)", "Group B (35%)", "Group C (25%)"]
X_tr, X_te, y_tr, y_te, g_tr, g_te = train_test_split(
X_base, y_base, groups, test_size=0.25, random_state=42, stratify=y_base
)
scaler_sg = StandardScaler()
X_tr_s = scaler_sg.fit_transform(X_tr)
model_sg = LogisticRegression(class_weight='balanced', random_state=42, max_iter=1000)
model_sg.fit(X_tr_s, y_tr)
subgroup_performance_analysis(model_sg, X_te, y_te, g_te, group_names,
scaler=scaler_sg, model_name="Diagnostic AI")Complete Medical AI Evaluation Summary Table
| Metric | Formula | Clinical Question Answered | When It Matters Most |
|---|---|---|---|
| Sensitivity | TP / (TP+FN) | Of all sick patients, how many did I catch? | Life-threatening disease; epidemic screening |
| Specificity | TN / (TN+FP) | Of all healthy patients, how many did I clear? | Rare disease; high-cost/risky follow-up |
| PPV | TP / (TP+FP) | If test positive, how likely is disease? | High-stakes treatment decisions |
| NPV | TN / (TN+FN) | If test negative, how likely is health? | Ruling out disease |
| LR+ | Sens / (1-Spec) | How much does a + result increase disease odds? | Bayesian updating of clinical probability |
| LR− | (1-Sens) / Spec | How much does a − result decrease disease odds? | Bayesian updating of clinical probability |
| AUC-ROC | Area under ROC | Overall discrimination ability | Comparing models; threshold-free comparison |
Common Mistakes in Medical AI Evaluation
Mistake 1: Reporting accuracy instead of sensitivity and specificity In clinical AI, accuracy is almost never the right primary metric. Always report sensitivity and specificity separately, because they measure fundamentally different things.
Mistake 2: Ignoring prevalence when interpreting PPV A PPV of 70% sounds reassuring — but if prevalence is 0.1%, that same test actually has a PPV closer to 2%. Always compute PPV with the actual prevalence of your target population.
Mistake 3: Validating only on hospital-referred populations Patients referred to specialist care tend to be sicker than the general population (verification bias or spectrum bias). A model validated on hospital referrals will overestimate its performance in a general screening population.
Mistake 4: Not reporting confidence intervals A sensitivity of “90%” from a 50-patient study is a very wide estimate. A 50-patient study provides an approximate 95% CI of [78%, 97%]. Clinical decision-making requires uncertainty quantification.
Mistake 5: Ignoring subgroup disparities A model that performs well on average may perform poorly on elderly patients, minority ethnic groups, or patients with comorbidities. Regulatory bodies now explicitly require subgroup analysis.
Mistake 6: Conflating sensitivity/specificity with PPV/NPV Sensitivity and specificity are properties of the test. PPV and NPV are properties of the test applied to a specific population. Presenting PPV figures from a hospital study to justify screening in a low-prevalence general population is a common and serious error.
Summary
Sensitivity and specificity are the foundational metrics of medical AI evaluation, and their meaning goes far deeper than their formulas suggest.
Sensitivity measures how completely a model captures real disease cases — its recall. High sensitivity is the price of safety in screening applications. Every percentage point of sensitivity you sacrifice translates directly to patients with real disease who are told they are healthy and receive no treatment.
Specificity measures how accurately a model clears healthy patients — its ability to avoid false alarms. High specificity is the price of efficiency and trust. Every percentage point of specificity you sacrifice translates to healthy people undergoing unnecessary, potentially harmful procedures.
The tradeoff between them — mediated by the decision threshold — is one of the most consequential decisions in medical AI deployment. Bayes’ theorem reveals that even excellent tests generate mostly false positives in low-prevalence populations, fundamentally changing how positive test results should be interpreted. Likelihood ratios provide the most portable way to quantify the evidential value of a test result across different prevalence settings.
For any medical AI system, real-world deployment requires sensitivity-specificity analysis across demographic subgroups, confidence intervals on all estimates, and calibration of the decision threshold to the specific clinical context — whether that is emergency screening (maximize sensitivity), confirmatory diagnosis (maximize specificity), or routine population screening (balance guided by the costs of each error type).